How to use viewmastR with large query objects
2024-12-10
BigQuery.RmdInstalling Rust
First you need to have an updated Rust installation. Go to this site to learn how to install Rust.
Installing viewmastR
You will need to have the devtools package installed…
devtools::install_github("furlan-lab/viewmastR")Running viewmastR
suppressPackageStartupMessages({
library(viewmastR)
library(Seurat)
library(ggplot2)
})
#query dataset
#seu <- readRDS(file.path(ROOT_DIR1, "240813_final_object.RDS"))
seu <- qs2::qs_read("/Users/sfurlan/Library/CloudStorage/OneDrive-SharedLibraries-FredHutchCancerCenter/Furlan_Lab_2025 - General/experiments/AML_MRD/DL2_SF/cds/260113_Filtered_Processed_vmAnnotated_scrublet.qs")
#reference dataset
seur<-readRDS(file.path(ROOT_DIR2, "230329_rnaAugmented_seurat.RDS"))
vg <- get_selected_features(seu)
vg <- vg[vg %in% rownames(seur)]Build the model and infer for small dataset (not using chunks and parallelization)
This is also covered elsewhere
results <- viewmastR(seu, seur, ref_celldata_col = "SFClassification", selected_features = vg, max_epochs = 4, train_only = T)
seu<-viewmastR_infer(seu, results[["model_dir"]], vg, labels = levels(factor(seur$SFClassification)))
DimPlot(seu, group.by = "viewmastR_inferred", cols = seur@misc$colors)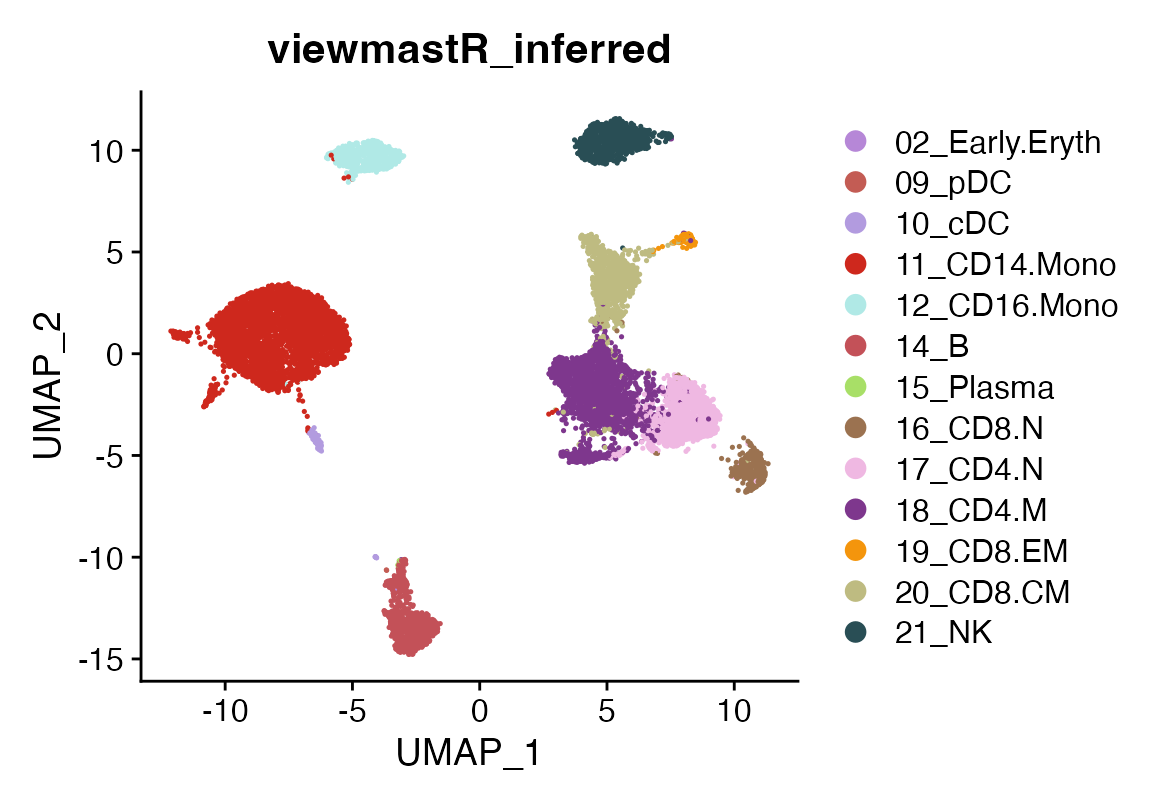
Build the model and infer for large dataset (dividing the query into chunks and using parallelization)
By using chunks and workers, you can infer from the model only chunks at a time using multiple workers in parallel.
gc()## used (Mb) gc trigger (Mb) limit (Mb) max used (Mb)
## Ncells 11296441 603.3 57654596 3079.1 NA 72068244 3848.9
## Vcells 772235464 5891.7 4742042616 36179.0 131072 8120631284 61955.5
system.time({
seu <- viewmastR_infer(seu, results[["model_dir"]],
query_celldata_col = "test1", vg,
labels = levels(factor(seur$SFClassification)),
threads = 1)
})## user system elapsed
## 132.740 45.308 183.008
# Parallel nd (8 threads)
system.time({
seu <- viewmastR_infer(seu, results[["model_dir"]],
query_celldata_col = "test2", vg,
labels = levels(factor(seur$SFClassification)),
threads = 8)
})## user system elapsed
## 144.396 59.958 168.579
all(seu$test1==seu$test2)## [1] TRUEWe see no difference if parallelization is used
confusion_matrix(pred = factor(seu$test1), gt = factor(seu$test2), cols = seur@misc$colors)